library(pxweb)
library(data.table)
library(ggplot2)
url <- "https://api.scb.se/OV0104/v1/doris/sv/ssd/BE/BE0101/BE0101A/BefolkningR1860N"
# PXWEB query
pxweb_query_list <-
list(
"Alder" = "*",
"Kon" = c("1", "2"),
"ContentsCode" = c("0000053A"),
"Tid" = as.character(1864:2024)
)
px_data <-
pxweb_get(
url = url,
query = pxweb_query_list
)
# A data.table with population numbers
bef <-
px_data |>
as.data.frame() |>
setDT()
# Some data cleaning
setnames(
bef,
c("ålder", "kön", "år", "Antal"),
c("age", "sex", "year", "N")
)
bef[, `:=`(
age = as.numeric(gsub(" år", "", age)),
sex = factor(sex, c("män", "kvinnor"), c("males", "females")),
year = as.integer(year)
)]
bef <- bef[!is.na(age)] # Remove totals
# Aggregate for age groups
bef[,
age_group := cut(
age,
c(-Inf, 17, 66, Inf),
c("children", "adults", "elderly")
)
]
bef_ag <- bef[, .(N = sum(N)), .(age_group, sex, year)]
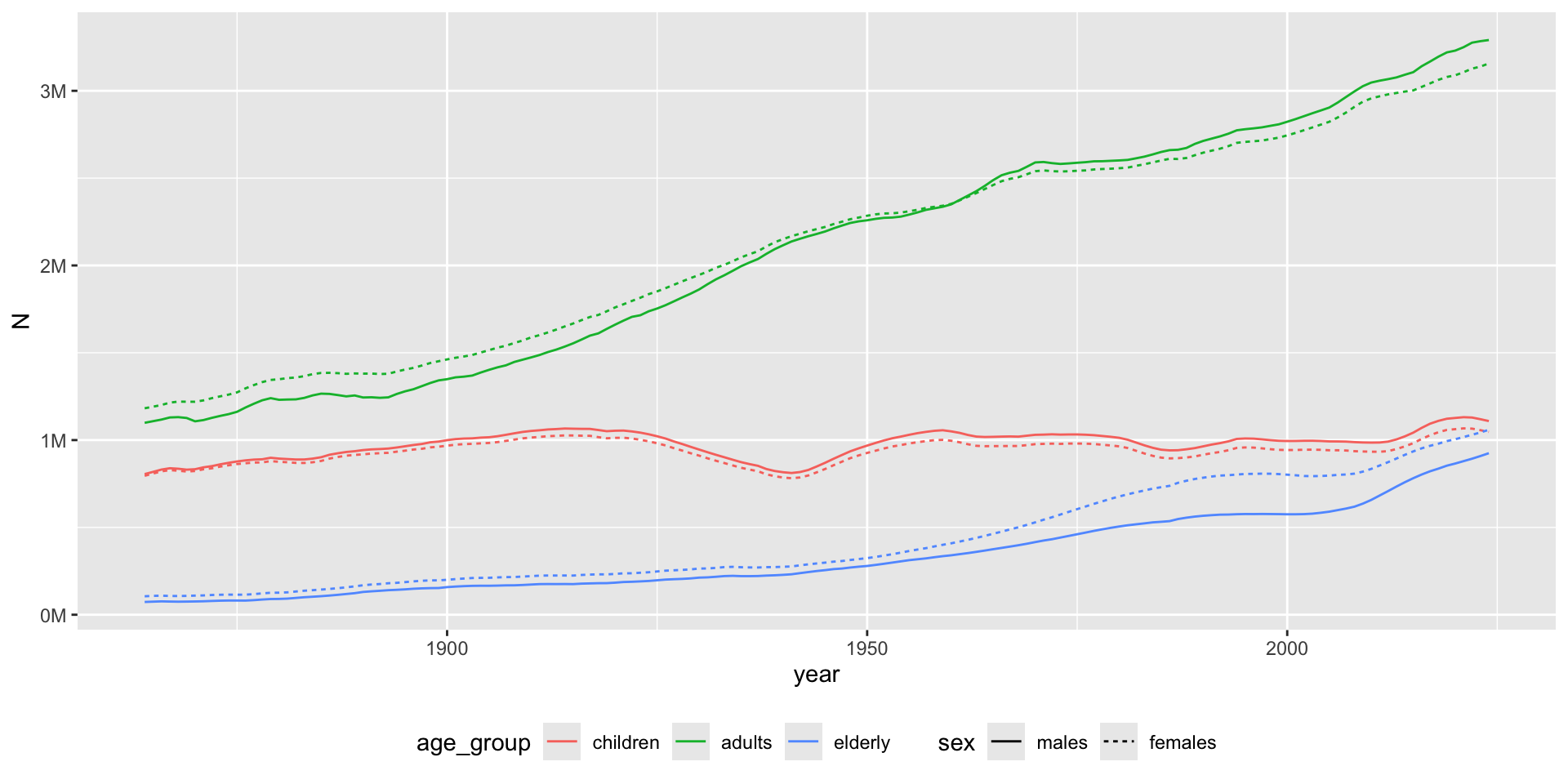
# Visualize
gg <- bef_ag |>
ggplot(aes(year, N, color = age_group, linetype = sex)) +
geom_line() +
theme(legend.position = "bottom") +
scale_y_continuous(
labels = scales::label_number(scale = 1e-6, suffix = "M")
)