###########################################################################
# #
# Purpose: My nice script #
# #
# Author: Erik Bülow #
# Contact: xxx@yyy.zzz #
# Client: John Doe #
# #
# Code created: 2026-03-20 #
# Last updated: 2026-03-20 #
# Source: /Users/xbuler/STA220/documentation #
# #
# Comment: Bla bla bla ... #
# #
###########################################################################EL9: Documentation
Quarto
Reading
- (Wickham, Çetinkaya-Rundel, and Grolemund, n.d., ch. 28-29) introduces Quarto with good examples
For reference
Levels of documentation
Documentation can exist at several levels in a project.
- Code level
- scripts (comments, e.g.
# clean data and merge cohorts) - functions (Roxygen2 documentation)
- notes and reminders (e.g.
# TODO:)
- scripts (comments, e.g.
- Project level
README.md- (GitHub repos may also have a wiki or a polished website)
- Output level
- figures and tables
- reports (PDF / HTML)
- presentations (slides)
Code level
Documenting R scripts
- Use Air for better formatting of the R code itself (will format the code according to best practice for readability and collaboration)
- Section labels by
Shift + Ctrl/Cmd + Rin Positron (and RStudio) - Easiest way to aid navigation to R scripts
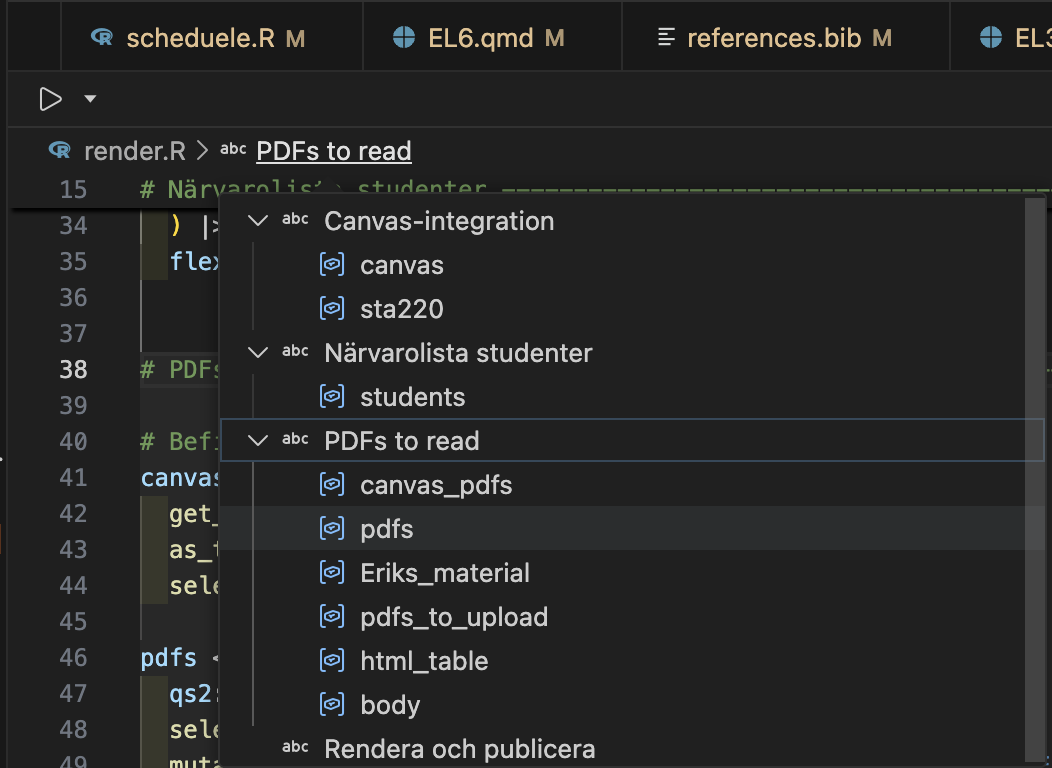
Individual scripts
- As soon as you distribute an individual R script from a bigger project (by e-mail or as a printed copy) some context would get lost.
- The commentr package is designed to maintain such context
remotes::install_bitbucket("cancercentrum/commentr")
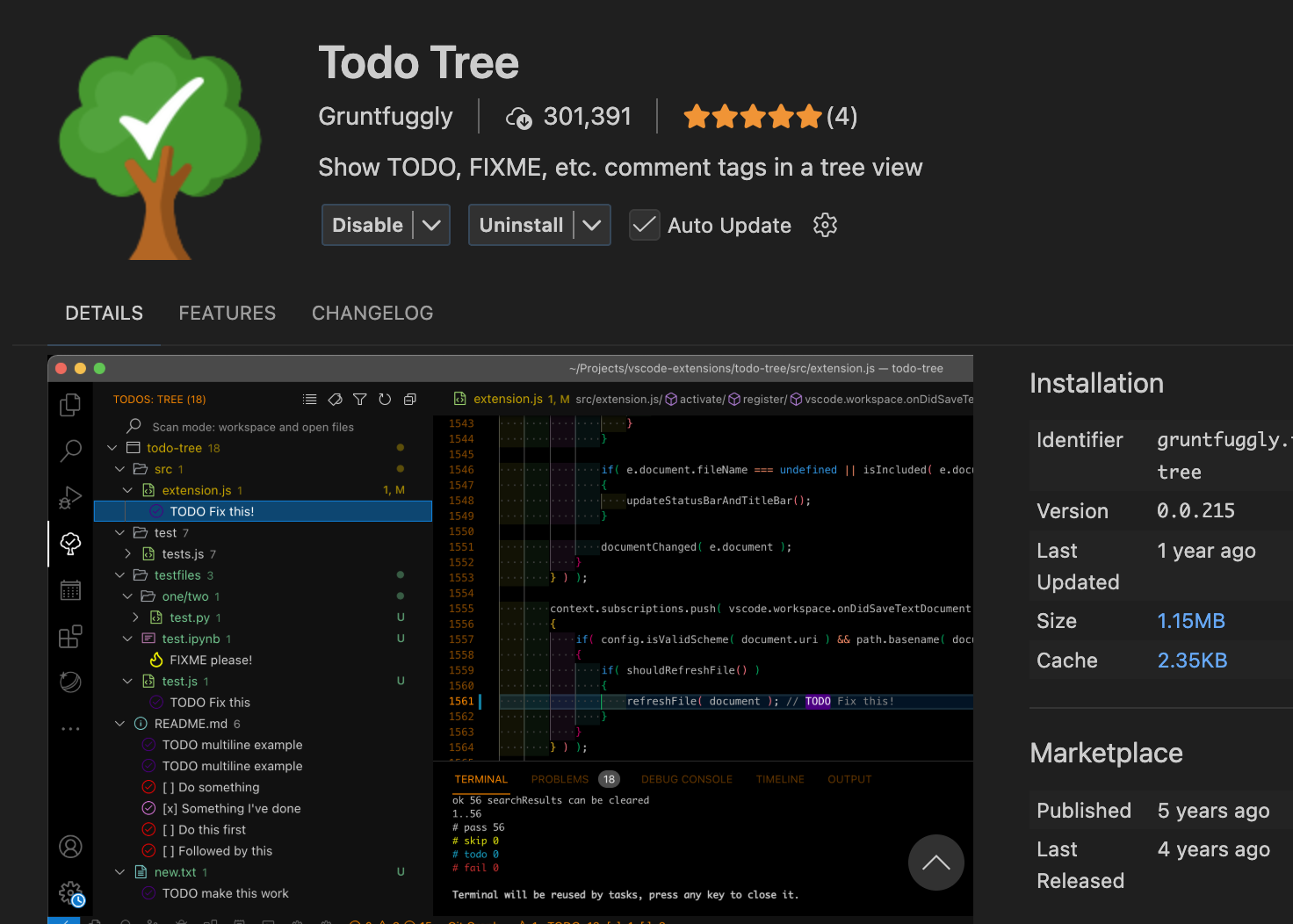
# TODO: Change this to something better:
1 + 2Documenting R functions
What does this function do?
#' Say hello
#'
#' Create a simple greeting. If no name is provided, the function
#' returns a greeting to the world.
#'
#' @param who A character vector giving the name(s) of the person or
#' object to greet. If missing, the greeting defaults to `"world"`.
#'
#' @return A character vector of the same length as `who` containing
#' greeting messages.
#'
#' @examples
#' hello()
#' hello("you")
#' hello(c("Alice", "Bob"))
hello <- function(who = "world") {
sprintf("Hello %s!", who)
}- There is a standardized syntax to document R functions
- You should be able to click a small light bulb (💡) to insert a sceleton when writing your function
- The same syntax is used within modern R packages
- but it is useful even within your own projects
- roxygen2 is a package used for package documentation but it also has a great manual for documenteing functions in general
Project level
Documenting a project
- You have a
README.mdfile within your git repository - It can contain plain text without any formatting
- It will be displayed up-front in GitHub
- You can also include formatting!
- It will be nicely rendered on GitHub or elsewhere
Basic formatting
| What you want | Markdown syntax | Result |
|---|---|---|
| Heading | # Heading |
Heading |
| Italic | *text* |
text |
| Bold | **text** |
text |
| Inline code | `code` |
code |
| Link | [text](https://example.com) |
text |
| Bullet list | - item |
• item |
| Numbered list | 1. item |
1. item |
Comment or heading
# is used for comments in R files and within R blocks but for headings in .md and .qmd files!
Visual mode
Instead of editing the source code, you may also use the “visual editor” in Positron:
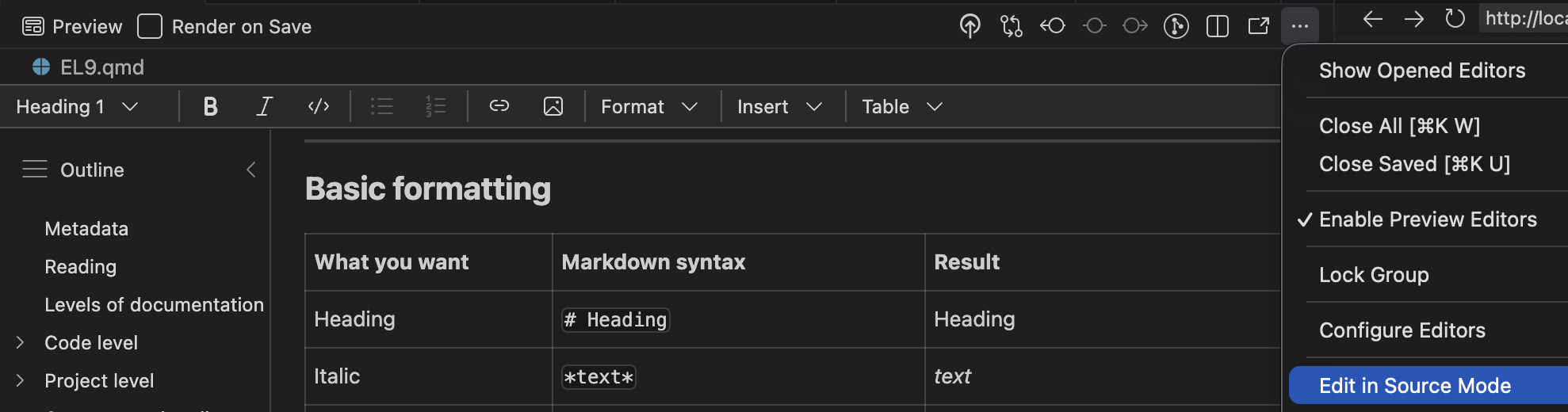
Warning
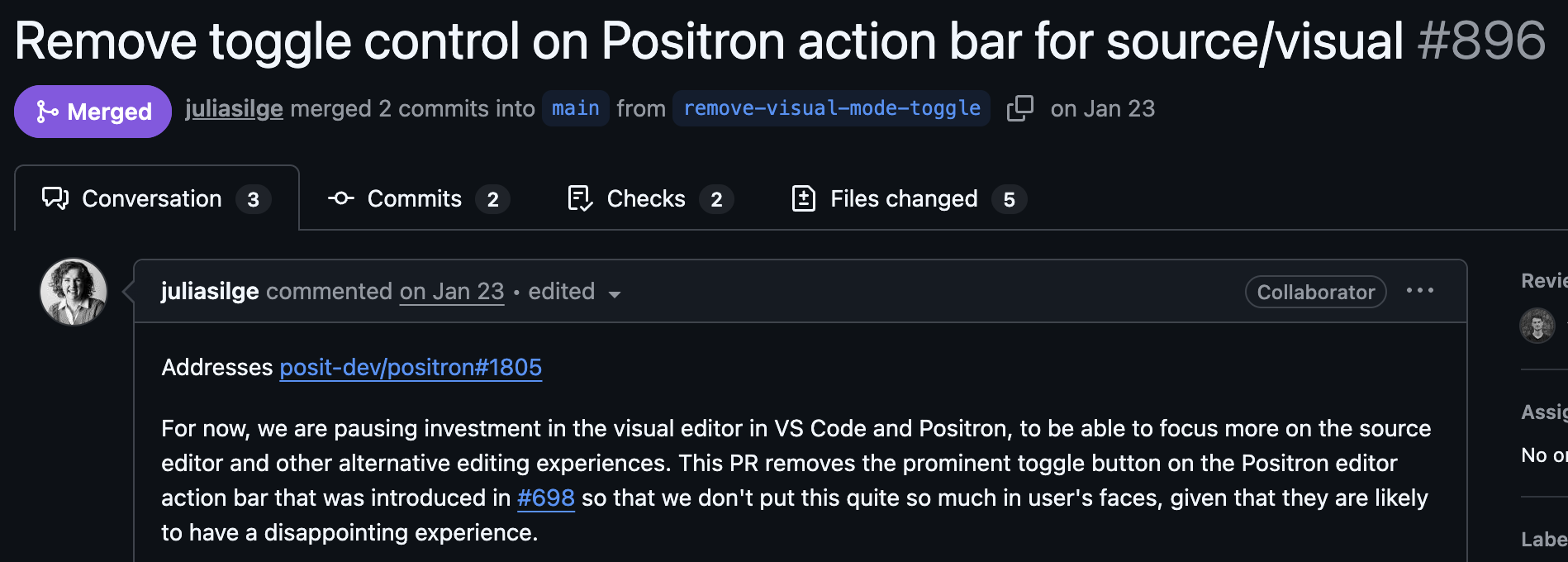
Use both?
Personally, I do most editing in source mode (better for the R chunks), but I find it much easier to include references/citations and to paste in external figures in using the visual mode.
Reproducible Workflows
Modern biostatistics projects rarely involve only statistics.
- data cleaning and preprocessing
- statistical modelling
- visualisation
- interpretation of results
- reporting and documentation
A good workflow should make it possible to combine all of these steps in one place.
Output level
What?
Quarto is a publishing system for scientific and technical documents.
It allows you to combine:
- text
- R code
- statistical results
- tables and figures
in a single workflow.
This makes it easier to create reproducible statistical reports.
Why?
In many projects people work with:
- R scripts for analysis 🥳
- Word documents for reporting 😱
- PowerPoint slides for presentations 😱
This separation often leads to:
- copy–paste errors 🥴
- outdated figures 👎
- results that cannot easily be reproduced 🤯
Quarto helps avoid these problems!
Reproducible analysis
In a Quarto document you can:
- write the explanation of your analysis
- include the R code used
- generate figures and tables automatically
- update everything by re-running the document
- If the data or model changes, the entire report can be regenerated automatically.
- It is widely used in data science, epidemiology and biostatistics.
- Integrated in Positron and developed by Posit PBC
Used in three ways
- For communicating to decision-makers, who want to focus on the conclusions, not the code behind the analysis.
- For collaborating with collegues (including future you!), who are interested in both your conclusions, and how you reached them (i.e. the code).
- As an environment in which to do data science and statistics, as a modern-day lab notebook where you can capture not only what you did, but also what you were thinking.
A simple report
When you chose New > New File ... > Quarto document, the file will contain:
- This is a YAML header (Yet Another Markup Language)
- Different output formats are available:
- html: can also be published online (Quartopub or GitHub pages)
- docx: Word document for collaborating with non-statisticians etc
- typst: PDF rendering (ever heard of \(\LaTeX\)? … This is the modern alternative!)
- revealjs: slide decks (you are looking at one!)
- Can render to multiple formats at once!
This is the yaml header for this presentation:
---
title: "EL9: Documentation"
subtitle: "Quarto"
reading: "[@wickham, ch. 28-29]"
format:
revealjs:
theme: blood
transition: fade
slide-number: true
css: styles.css
smaller: true
scrollable: true
incremental: false
typst:
include-before-body:
- text: |
// Gör alla länkar tydliga i PDF: blå + understrukna
#show link: set text(fill: rgb("1a5fb4"))
#show link: underline
docx: default
bibliography: references.bib
---YAML structure
YAML uses indentation to represent structure.
Think of it like a tree.
This corresponds to the structure:
format
└── html
└── code-fold: trueIf indentation is wrong?
Now Quarto interprets this as:
format
html
└── code-fold: trueThis is not the intended structure, and the document may fail to render.
💡 Rule of thumb
- use two spaces per level
- never use tabs
- when something fails, check indentation first
Three parts
Quarto documents have three parts:
- An (optional) YAML header surrounded by
---s. - Chunks of R code surrounded by
```{r}and```.
Keybindings: Cmd + option + I (Mac) or Ctrl + Alt + I (Windows)
- Text with markdown format
Order
Part 1 (YAML header) always comes first but 2 (code block) and 3 (Markdown text) are usually intermingled throughout the rest of the document.
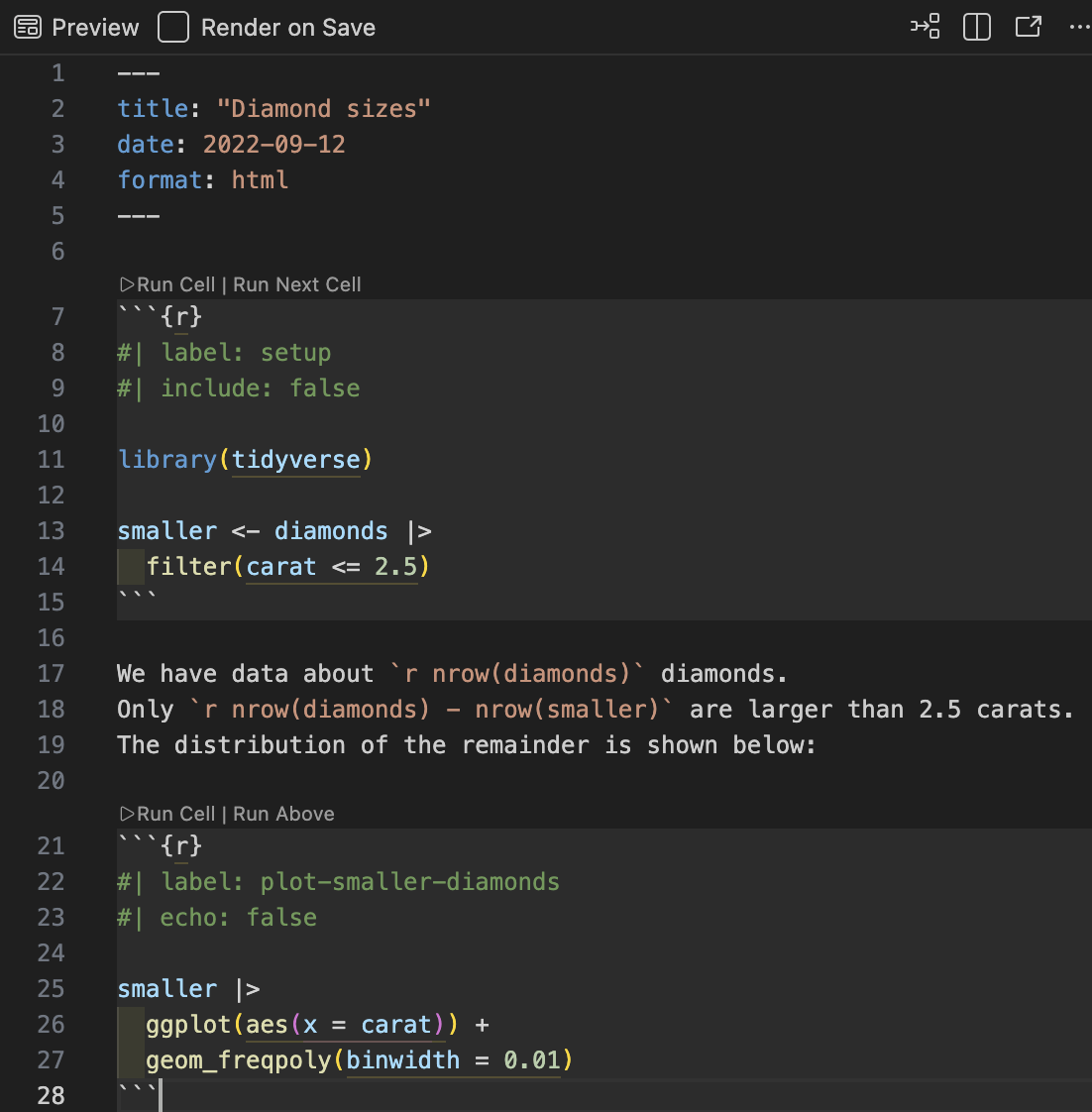
Inline code
Sometimes we want to insert small pieces of R output directly in the text.
This is called inline code.
Inline code will always just show the result of the call (not the code itself in the rendered document):
Todys date is 2026-03-20 and it is 12 PM
To higlight with bold and italic:
Todays date is 2026-03-20 and it is 12 PM
To publish
Decision makers, stake holders, friends or the public might not care about R. Disable R code (echoing the code itself), warnings and messages:
For collaborators
- Might still care less about verbose R messages and warnings (they trust that you have already considered those)
- May still need the underlaying R code for deeper understaing
- Use
code-fold
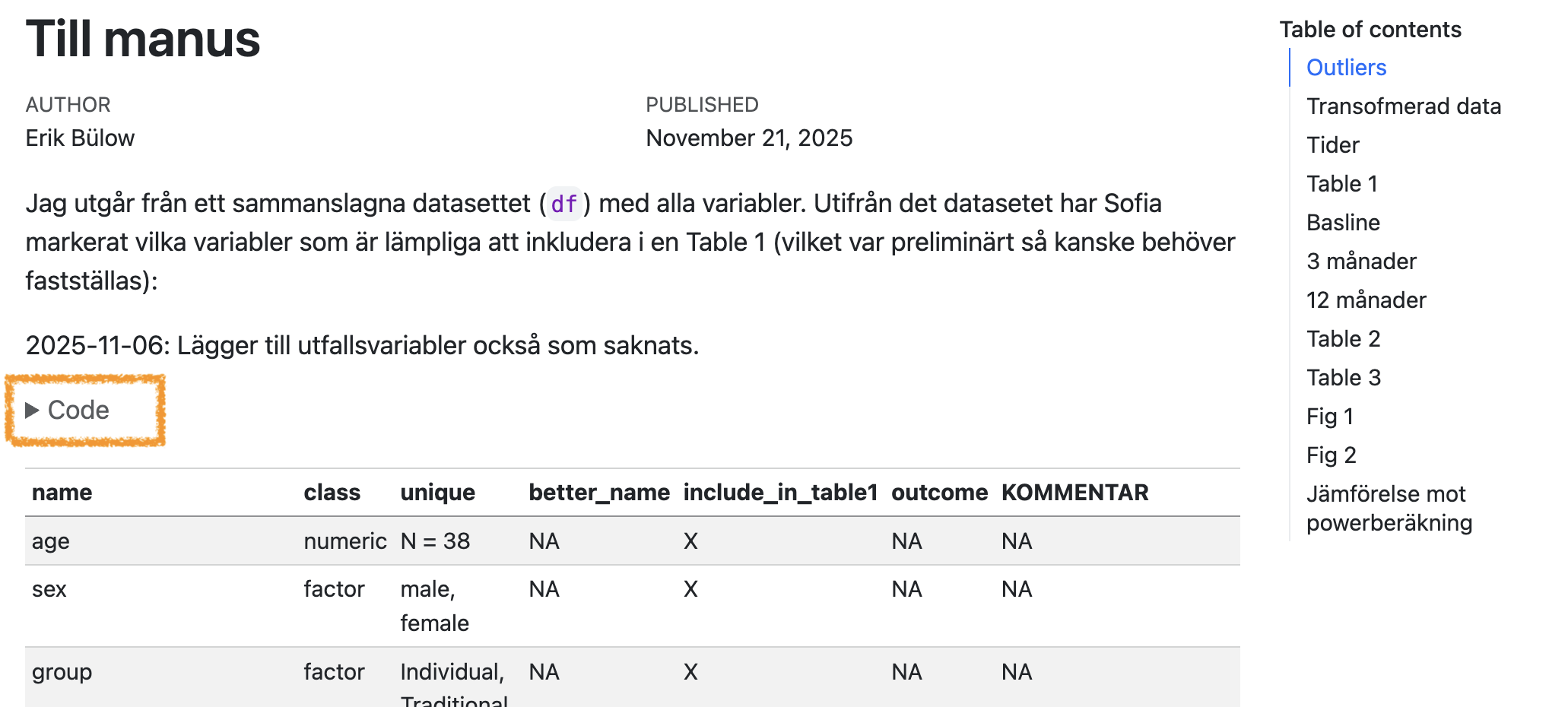
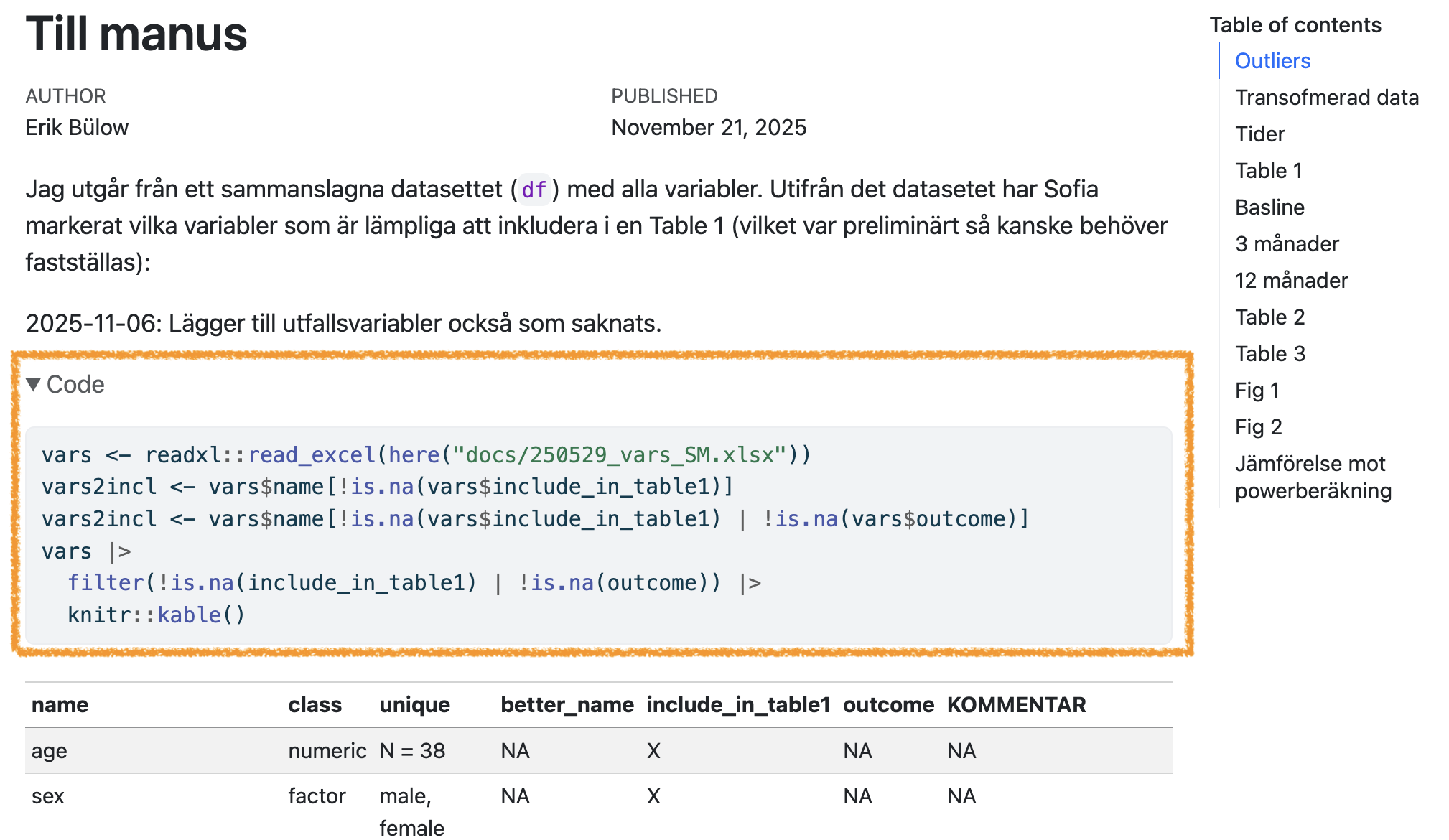
This can also be set individually for each code chunk:
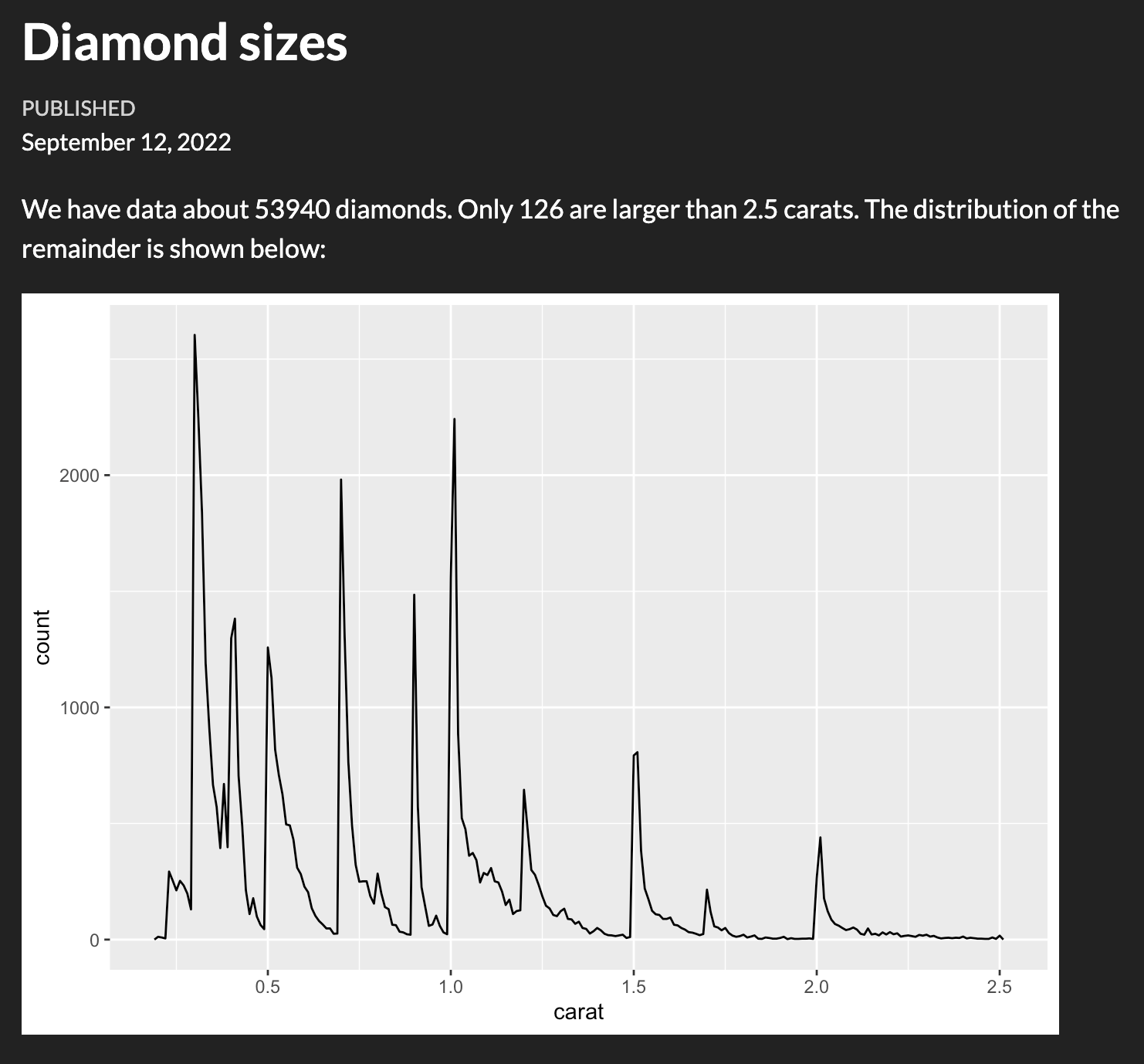
Figures
Figures are usualy displayed without problem (and you can adjust their size, add captions etc):
This is origo!
Tables
Just printing the R output might not look very nice:
Better with kable
The easiest way to make tibbles/data.frames/data.tables look nicer is with df-print: kable in the yaml header.
Or manually for individual outputs:
| Sepal.Length | Sepal.Width | Petal.Length | Petal.Width | Species |
|---|---|---|---|---|
| 5.1 | 3.5 | 1.4 | 0.2 | setosa |
| 4.9 | 3.0 | 1.4 | 0.2 | setosa |
| 4.7 | 3.2 | 1.3 | 0.2 | setosa |
| 4.6 | 3.1 | 1.5 | 0.2 | setosa |
| 5.0 | 3.6 | 1.4 | 0.2 | setosa |
| 5.4 | 3.9 | 1.7 | 0.4 | setosa |
Best with gt
- Very flexible
- Publication ready
- Package from Posit
- flextable may be preferred instead if rendering Office documents (Word, PowerPoint)
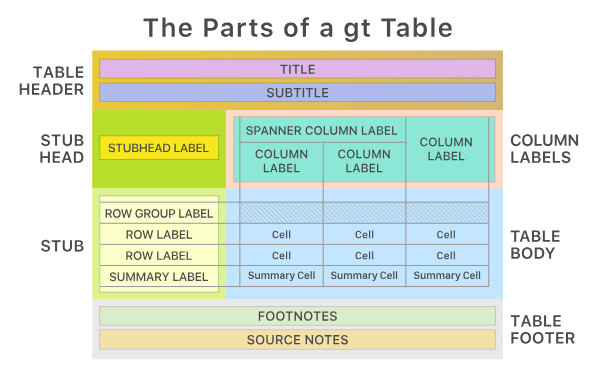
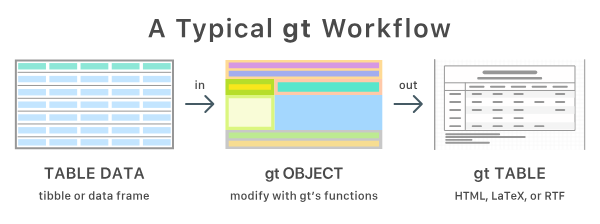
gtsummary
gtsummary package build on top of {gt} to easily summarise datasets, regression models etc
| Characteristic | setosa N = 501 |
versicolor N = 501 |
virginica N = 501 |
|---|---|---|---|
| Sepal.Length | 5.00 (4.80, 5.20) | 5.90 (5.60, 6.30) | 6.50 (6.20, 6.90) |
| Sepal.Width | 3.40 (3.20, 3.70) | 2.80 (2.50, 3.00) | 3.00 (2.80, 3.20) |
| Petal.Length | 1.50 (1.40, 1.60) | 4.35 (4.00, 4.60) | 5.55 (5.10, 5.90) |
| Petal.Width | 0.20 (0.20, 0.30) | 1.30 (1.20, 1.50) | 2.00 (1.80, 2.30) |
| 1 Median (Q1, Q3) | |||
Presentations
- Quarto can export to PowerPoint
- And to the older Beamer format
- But I recommend revealjs
format: revealjs- can also be modified with speaker notes, laser pointer etc
- Info and examples
For example (as used for these slides):
Integrate with {targets}
in _targets.R
In report.qmd
Try it
Copy this code to the terminal (not the console!) and try targets::tar_make() in the console!
cd
mkdir test_targets_quarto
cat > test_targets_quarto/_targets.R <<'EOF'
library(targets)
library(tarchetypes) # Lets you integrate a quarto report/presentation
list(
tar_target(data, iris),
tar_target(model, lm(Sepal.Length ~ ., data)),
tar_quarto(report, "report.qmd") # Make presentation
)
EOF
cat > test_targets_quarto/report.qmd <<'EOF'
---
title: My project
author: My name
date: today
format:
typst: default
docx: default
html: default
revealjs:
output-file: slides.html
---
## Here are some flowers!
gtsummary::tbl_summary(targets::tar_read(data))
## And here is a model
gtsummary::tbl_regression(targets::tar_read(model))
EOF
positron test_targets_quarto
# rm -r test_targets_quartoManuscripts
- Quarto Manuscript
- framework for writing and publishing scholarly articles
- Something for your later thesis work?
- Handles references
- Export to Word documents (
.docx) etc- which is still the most common format for submission to medical/epidemiological journals
- Still used for collaborative work with non-statisticians etc
Use references
When writing scientific reports we need to:
- acknowledge previous research
- support claims with evidence
- allow readers to find the original sources
This is done through citations and a reference list.
Example:
Don’t blame me, it was according to Wickham, Çetinkaya-Rundel, and Grolemund (n.d.).
At the end of the document a bibliography is automatically generated.
How Quarto handles references
Quarto uses a bibliography file, usually in .bib format.
Example in the document header:
Then citations can be written directly in the text:
Several studies have examined this relationship @smith2020.Quarto will render something like:
Several studies have examined this relationship (Smith 2020).
The full reference will appear automatically in the bibliography.
Online portfolio
- Looking for a job after you fininsh the master program?
- An online portfolia might increase your visability!
- Perhaps add your course project to such portfolio?
- Demo